Imputation as an approach to missing data has been around for decades.
You probably learned about mean imputation in methods classes, only to be told to never do it for a variety of very good reasons. Mean imputation, in which each missing value is replaced, or imputed, with the mean of observed values of that variable, is not the only type of imputation, however.
Two Criteria for Better Imputations
Better, although still problematic, imputation methods have two qualities. They use other variables in the data set to predict the missing value, and they contain a random component.
Using other variables preserves the relationships among variables in the imputations. It feels like cheating, but it isn’t. It ensures that any estimates of relationships using the imputed variable are not too low. Sure, underestimates are conservatively biased, but they’re still biased.
The random component is important so that all missing values of a single variable are not exactly equal. Why is that important? If all imputed values are equal, standard errors for statistics using that variable will be artificially low.
There are a few different ways to meet these criteria. One example would be to use a regression equation to predict missing values, then add a random error term.
Although this approach solves many of the problems inherent in mean imputation, one problem remains. Because the imputed value is an estimate–a predicted value–there is uncertainty about its true value. Every statistic has uncertainty, measured by its standard error. Statistics computed using imputed data have even more uncertainty than their standard errors measure.
Your statistical package cannot distinguish between an imputed value and a real value.
Since the standard errors of statistics based on imputed values are too small, corresponding p-values are also too small. P-values that are reported as smaller than they actually are? Those lead to Type I errors.
How Multiple Imputation Works
Multiple imputation solves this problem by incorporating the uncertainty inherent in imputation. It has four 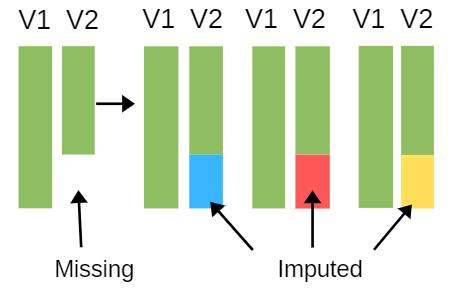 steps:
steps:
- Create m sets of imputations for the missing values using a good imputation process. This means it uses information from other variables and has a random component.
- The result is m full data sets. Each data set will have slightly different values for the imputed data because of the random component.
- Analyze each completed data set. Each set of parameter estimates will differ slightly because the data differs slightly.
- Combine results, calculating the variation in parameter estimates.
Remarkably, m, the number of sufficient imputations, can be only 5 to 10 imputations, although it depends on the percentage of data that are missing. A good multiple imputation model results in unbiased parameter estimates and a full sample size.
Doing multiple imputation well, however, is not always quick or easy. First, it requires that the missing data be missing at random. Second, it requires a very good imputation model. Creating a good imputation model requires knowing your data very well and having variables that will predict missing values.
Updated 9/20/2021



Thanks for the neat summary of multiple imputation. It’s also available in R, e.g., https://cran.r-project.org/web/packages/mice/index.html