I’m sure you’ve heard that R creates beautiful graphics.
It’s true, and it doesn’t have to be hard to do so. Let’s start with a simple histogram using the hist() command, which is easy to use, but actually quite sophisticated.
First, we set up a vector of numbers and then we create a histogram.
B <- c(2, 4, 5, 7, 12, 14, 16)
hist(B)
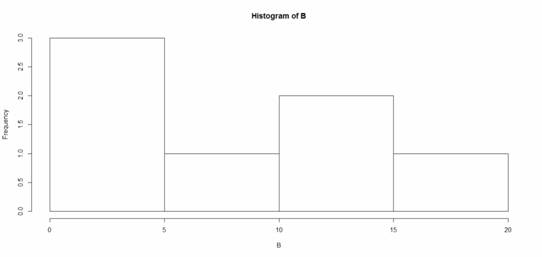
That was easy, but you need more from your histogram. Note that R decided on an appropriate bin width. OK. Now we create a histogram from all the data in an array.
A <- structure(list(James = c(1L, 3L, 6L, 4L, 9L), Robert = c(2L, 5L,
4L, 5L, 12L), David = c(4L, 4L, 6L, 6L, 16L), Anne = c(3L, 5L,
6L, 7L, 6L)), .Names = c("James", "Robert", "David", "Anne"), class = "data.frame", row.names = c(NA, -5L))
attach(A)
A
James Robert David Anne
1 1 2 4 3
2 3 5 4 5
3 6 4 6 6
4 4 5 6 7
5 9 12 16 6
The trick is to transform the four variables into a single vector and make a histogram of all elements.
B <- c(A$James, A$Robert, A$David, A$Anne)
Let’s create a histogram of B in dark green and include axis labels.
hist(B, col="darkgreen", ylim=c(0,10), ylab ="MY HISTOGRAM", xlab
="FREQUENCY")
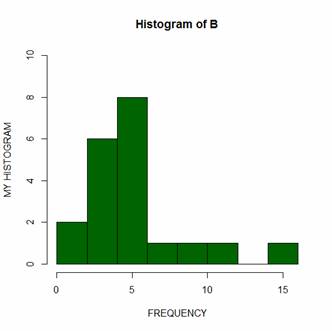
However, controlling bin numbers can be problematic. Try setting the number of bins at 6 using the breaks argument.
hist(B, col = "red", breaks=6, ylim=c(0,8),
main="My Histogram", las=2, xlab = "Values", cex.lab = 1.3)
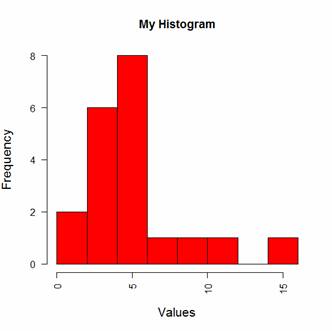
You can see that R has taken the number of bins (6) as indicative only. However, setting up histogram bins as a vector gives you more control over the output. Now we set up the bins as a vector, each bin four units wide, and starting at zero.
bins<- c(0, 4, 8, 12, 16)
hist(B, col = "blue", breaks=bins, ylim=c(0,8),
main="My Histogram", las=2, xlab = "Values", cex.lab = 1.3)
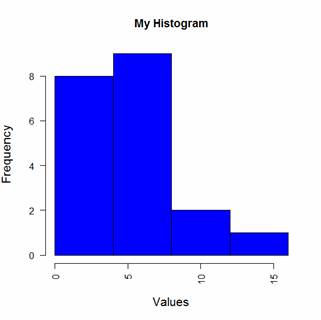
Now we have four bins of the right width. That wasn’t so hard! In Part 13 we will look at further plotting techniques in R.
About the Author: David Lillis has taught R to many researchers and statisticians. His company, Sigma Statistics and Research Limited, provides both on-line instruction and face-to-face workshops on R, and coding services in R. David holds a doctorate in applied statistics.
See our full R Tutorial Series and other blog posts regarding R programming.

Again, the table does not read in right using “attach”–maybe it’s due to an R update, or to being on a Mac. But given that the prior one didn’t read in correctly even after jumping through hoops… is there an update needed, or something mac specific?
Here’s what copy-paste gives; it looks like it might be a problem with the tab (or spaces) between the first two columns, which might be handled differently using a mac?:
> A
James Robert David Anne
1 1 2 4 3
2 3 5 4 5
3 6 4 6 6
4 4 5 6 7
5 9 12 16 6
> James Robert David Anne
Error: unexpected symbol in ” James Robert”
> 1 1 2 4 3
Error: unexpected numeric constant in “1 1”
> 2 3 5 4 5
Error: unexpected numeric constant in “2 3”
> 3 6 4 6 6
Error: unexpected numeric constant in “3 6”
> 4 4 5 6 7
Error: unexpected numeric constant in “4 4”
> 5 9 12 16 6
Error: unexpected numeric constant in “5 9”
R may not be a good choice for sercah engine optimisation, but they’ve overcome it, I sercahed for R on Google and it was the first result.I’m frustrated with normal stats programs (in some respects, in others I love them), and I’m doing a 3 day R course to see if it can scratch my statistical itch. I’m looking forwards to it, the last time I used a programming language was BASIC in the mid 90s.