One important yet difficult skill in statistics is choosing a type model for different data situations. One key consideration is the dependent variable.
For linear models, the dependent variable doesn’t have to be normally distributed, but it does have to be continuous, unbounded, and measured on an interval or ratio scale.
Percentages don’t fit these criteria. Yes, they’re continuous and ratio scale. The issue is the (more…)
Interpreting regression coefficients can be tricky, especially when the model has interactions or categorical predictors (or worse – both).
But there is a secret weapon that can help you make sense of your regression results: marginal means.
They’re not the same as descriptive stats. They aren’t usually included by default in our output. And they sometimes go by the name LS or Least-Square means.
And they’re your new best friend.
So what are these mysterious, helpful creatures?
What do they tell us, really? And how can we use them?
(more…)
One question that seems to come up pretty often is:
What is the difference between logistic and probit regression?
Well, let’s start with how they’re the same:
Both are types of generalized linear models. This means they have this form:
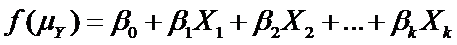
(more…)
An incredibly useful tool in evaluating and comparing predictive models is the ROC curve.
Its name is indeed strange. ROC stands for Receiver Operating Characteristic. Its origin is from sonar back in the 1940s. ROCs were used to measure how well a sonar signal (e.g., from an enemy submarine) could be detected from noise (a school of fish).
ROC curves are a nice way to see how any predictive model can distinguish between the true positives and negatives. (more…)
When you have data measuring the time to an event, you can examine the relationship between various predictor variables and the time to the event using a Cox proportional hazards model.

In this webinar, you will see what a hazard function is and describe the interpretations of increasing, decreasing, and constant hazard. Then you will examine the log rank test, a simple test closely tied to the Kaplan-Meier curve, and the Cox proportional hazards model.
Note: This training is an exclusive benefit to members of the Statistically Speaking Membership Program and part of the Stat’s Amore Trainings Series. Each Stat’s Amore Training is approximately 90 minutes long.
(more…)
I received a question recently about R Commander, a free R package.
R Commander overlays a menu-based interface to R, so just like SPSS or JMP, you can run analyses using menus. Nice, huh?
The question was whether R Commander does everything R does, or just a small subset.
Unfortunately, R Commander can’t do everything R does. Not even close.
But it does a lot. More than just the basics.
So I thought I would show you some of the things R Commander can do entirely through menus–no programming required, just so you can see just how unbelievably useful it is.
Since R commander is a free R package, it can be installed easily through R! Just type install.packages("Rcmdr") in the command line the first time you use it, then type library("Rcmdr") each time you want to launch the menus.
Data Sets and Variables
Import data sets from other software:
- SPSS
- Stata
- Excel
- Minitab
- Text
- SAS Xport
Define Numerical Variables as categorical and label the values
Open the data sets that come with R packages
Merge Data Sets
Edit and show the data in a data spreadsheet
Personally, I think that if this was all R Commander did, it would be incredibly useful. These are the types of things I just cannot remember all the commands for, since I just don’t use R often enough.
Data Analysis
Yes, R Commander does many of the simple statistical tests you’d expect:
- Chi-square tests
- Paired and Independent Samples t-tests
- Tests of Proportions
- Common nonparametrics, like Friedman, Wilcoxon, and Kruskal-Wallis tests
- One-way ANOVA and simple linear regression
What is surprising though, is how many higher-level statistics and models it runs:
- Hierarchical and K-Means Cluster analysis (with 7 linkage methods and 4 options of distance measures)
- Principal Components and Factor Analysis
- Linear Regression (with model selection, influence statistics, and multicollinearity diagnostic options, among others)
- Logistic regression for binary, ordinal, and multinomial responses
- Generalized linear models, including Gamma and Poisson models
In other words–you can use R Commander to run in R most of the analyses that most researchers need.
Graphs
A sample of the types of graphs R Commander creates in R without you having to write any code:
- QQ Plots
- Scatter plots
- Histograms
- Box Plots
- Bar Charts
The nice part is that it does not only do simple versions of these plots. You can, for example, add regression lines to a scatter plot or run histograms by a grouping factor.
If you’re ready to get started practicing, click here to learn about making scatterplots in R commander, or click here to learn how to use R commander to sample from a uniform distribution.



